Difference between revisions of "LipidXplorer msconvert conversion"
(→Download and installation) |
(→Download and installation) |
||
| (2 intermediate revisions by the same user not shown) | |||
| Line 17: | Line 17: | ||
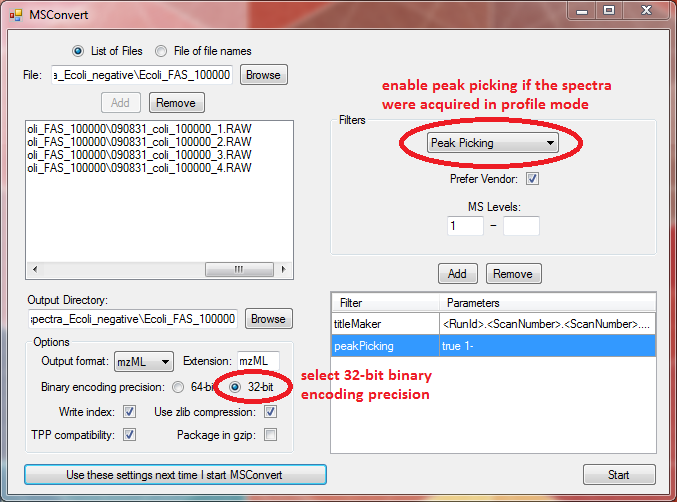
For LipidXplorer set the options in MSConvert like this: | For LipidXplorer set the options in MSConvert like this: | ||
| − | [[Image: | + | [[Image:MSConvertSettings.png|center||LipidXplorer Workflow]] |
For convenience store your settings by clicking "Use these settings next time I start MSConvert". Then upload your file(s) using "Browse" and "Add", respectively. | For convenience store your settings by clicking "Use these settings next time I start MSConvert". Then upload your file(s) using "Browse" and "Add", respectively. | ||
| Line 23: | Line 23: | ||
Use "Start" to do the conversion. | Use "Start" to do the conversion. | ||
| − | + | Always select 32-bit binary encoding precision. | |
| + | |||
| + | '''Important:''' If you have your spectra imported in profile mode you have to convert it to a centroid spectrum by peak picking. To switch on the peak picking select it from the filters on the upper right. Select the MS level 1- and click "Add" (see screenshot above). Now the peak picker is enabled. | ||
Latest revision as of 14:54, 3 April 2013
MSConvert
Only a few mass spectrometers directly produce mzXML/mzML files but there are several tools available that generate mzXML/mzML files from native acquired files.
MSConvert is the currently most featured converter (MSConvert) and further it is under ongoing maintenance. It supports various vendor file formats:
- Agilent Mass Hunter Data Access Component Library
- Waters Raw Data Access Component Library
- Bruker CompassXtract
- Thermo-Scientific MSFileReader Library
- Waters Raw Data Access Component Library
- AB Sciex WIFF Reader Library
Download and installation
Download MSConvert from MSConvert. It is part of Protewizard, a tool for proteomics. Just click on the button "I agree to the license terms, download Proteowizard" and install the downloaded file.
For LipidXplorer set the options in MSConvert like this:
For convenience store your settings by clicking "Use these settings next time I start MSConvert". Then upload your file(s) using "Browse" and "Add", respectively.
Use "Start" to do the conversion.
Always select 32-bit binary encoding precision.
Important: If you have your spectra imported in profile mode you have to convert it to a centroid spectrum by peak picking. To switch on the peak picking select it from the filters on the upper right. Select the MS level 1- and click "Add" (see screenshot above). Now the peak picker is enabled.